Applications¶
A diverse set of common HPC and various commercial applications are made available to all users on both POD MT1 & MT2 clusters. Chances are good that the specific application/version is already installed and ready to use as part of your workflow on POD. Users can also bring their own pre-built applications and code to enable their workflow on POD. New versions of commercial and open source applications are released all the time and can be installed by the POD support team if required. If you need additional software installed, please email pod@penguincomputing.com.
Environment Modules¶
The POD MT1 & MT2 clusters use Environment Modules to make these applications available to users. Loading a module will updates your user environment to make a specific build or version available for your use. Please reference the documentation below for more information.
Show Pre-installed Applications¶
To display the list of all pre-installed applications use the module avail command. You can narrow down the list by specifying an application name. This example output is truncated because of the large number of application currently available. Running the command module avail fluent only displays the different version of the ANSYS Fluent application installed on MT1.
$ module avail
----------------------------- /public/modulefiles ------------------------------
atlas/3.10.1/gcc.4.4.7 OpenFoam/2.2.1
biopython/1.6.1/python.2.7.4 openmpi/1.2.9/intel.12.1.0
boost/1.53.0/gcc.4.4.7 openmpi/1.4.5/gcc.4.4.7
boto/2.9.4/python.2.7.4 openmpi/1.4.5/gcc.4.7.2
cherrypy/3.2.2/python.2.7.4 openmpi/1.4.5/intel.11.1.0
cmake/2.8.11.2 openmpi/1.4.5/intel.12.1.0
fftw/3.3.3/gcc.4.4.7 openmpi/1.5.5/gcc.4.4.7(default)
fluent/14.5 openmpi/1.5.5/gcc.4.7.2
fluent/15.0 openmpi/1.5.5/intel.11.1.0
gcc/4.4.7(default) openmpi/1.5.5/intel.12.1.0
gcc/4.7.2 openmpi/1.6.4/gcc.4.4.7
...
$ module avail fluent
----------------------------- /public/modulefiles ------------------------------
fluent/14.5 fluent/15.0
Load an Environment Module to Access an Application¶
All applications are loaded into your environment using the module load command. Loading an environment module will update your $PATH, $LD_LIBRARY_PATH and $MANPATH environment variables for the specific version and build of the application you chose. Additionally, any dependent modules will be loaded into your environment as well. For example, loading the OpenFoam/2.2.1 module on MT1 will also load the appropriate OpenMPI and GCC compiler modules. Use the module list command to display the list of currently loaded modules.
$ module load OpenFoam/2.2.1
$ module list
Currently Loaded Modulefiles:
1) gcc/4.7.2 3) OpenFoam/2.2.1
2) openmpi/1.6.4/gcc.4.7.2
Switching Between Applications Versions¶
If you need to change applications or use a different version of a specific application use the module purge command to clear all your loaded modules returning your environment to it’s default state. For example, after loading the OpenFoam/2.2.1 module you want to switch to OpenFoam/2.3.1 make sure to purge your environment before loading the new one. Observe that OpenFoam/2.3.1 users a different set of dependent modules.
$ module load OpenFoam/2.2.1
$ module list
Currently Loaded Modulefiles:
1) gcc/4.7.2 3) OpenFoam/2.2.1
2) openmpi/1.6.4/gcc.4.7.2
$ module purge
$ module list
No Modulefiles Currently Loaded.
$ module load OpenFoam/2.3.1
$ module list
Currently Loaded Modulefiles:
1) gcc/4.7.2 3) boost/1.53.0/gcc.4.7.2 5) qt/4.8.5 7) OpenFoam/2.3.1
2) openmpi/1.6.4/gcc.4.7.2 4) cmake/2.8.11.2 6) mesa/9.1.6/gcc.4.7.2
User Defined Modules¶
If you install software in your home directory and wish to create your own set of modules, update the $MODULEPATH environment variable to include the directory containing your custom set of modules. For example, to use your custom modules in the $HOME/modulefiles directory, add this line to your ~/.bashrc. You should see your custom modules listed in the module avail command output and can now load them in your PBS TORQUE scripts.
export MODULEPATH="$HOME/modulefiles:$MODULEPATH"
FEA, CFD and FDTD Modeling¶
LS-DYNA is an advanced general-purpose multiphysics simulation software package. It’s core-competency lies in highly nonlinear transient dynamic Finite Element Analysis (FEA) using explicit time integration.
LS-PREPOST is an advanced pre-/post-processing utility that is delivered free with LS-DYNA.
LS-OPT is an optimization and probabilistic analysis program that can interface with LS-DYNA.
OpenFOAM (Open source Field Operation And Manipulation) is a C++ toolbox for the development of customized numerical solvers, and pre-/post-processing utilities for the solution of continuum mechanics problems, including Computational Fluid Dynamics (CFD).
ANSYS HFSS is the industry standard for simulating 3-D full-wave electromagnetic fields. Its gold-standard accuracy, advanced solver and compute technology have made it an essential tool for engineers designing high-frequency and high-speed electronic components.
ANSYS Fluent contains the broad physical modeling capabilities needed to model flow, turbulence, heat transfer, and reactions for industrial applications.
StarCCM+ is CD-adapco´s newest CFD software product. It uses the well established CFD solver technologies available in STAR-CD, and it employs a new client-server architecture and object oriented user interface to provide a highly integrated and powerful CFD analysis environment to users.
CONVERGE is a Computational Fluid Dynamics (CFD) code that completely eliminates the user time needed to generate a mesh through an innovative run-time mesh generation technique.
Lumerical simulation tools implement FDTD algorithms.
SU2 an open source collection of C++ based software tools for performing Partial Differential Equation (PDE) analysis and solving PDE constrained optimization problems.
AVL FIRE is a powerful multi-purpose thermo-fluid software representing the latest generation of 3-D CFD.
DSS SIMULIA Abaqus Unified FEA product suite offers powerful and complete solutions for both routine and sophisticated engineering problems covering a vast spectrum of industrial applications.
Weather Modeling and Cartographic Projections¶
WRF (Weather Research and Forecasting) model is a numerical weather prediction system.
COAMPS® (Coupled Ocean/Atmosphere Mesoscale Prediction System), developed and run by the Naval Research Laboratory in Monterey, CA, is the numerical model used for wind nowcasts and forecasts.
GRIB2 (GRIdded Binary or General Regularly-distributed Information in Binary form) is a common data format commonly used to store weather data.
PROJ is a cartographic projections library.
Life Sciences, BioTech and Genomics¶
LAMMPS (Large-scale Atomic/Molecular Massively Parallel Simulator) is a molecular dynamics program that makes use of MPI for parallel communication.
SOAPdenovo (Short Oligonucleotide Analysis Package) is a bioinformatics package used for the assembly and analysis of DNA sequences.
ABySS (Assembly By Short Sequences) is a de novo, parallel, paired-end sequence assembler that is designed for short reads.
LifeScope is genomic analysis software for SOLiD next-generation sequencing
BioPerl is a collection of Perl modules that facilitate the development of Perl scripts for bioinformatics applications.
Bowtie is an ultrafast, memory-efficient short read aligner for short DNA sequences.
Burrows-Wheeler Aligner (BWA) is a software package for mapping low-divergent sequences against a large reference genome, such as the human genome.
Celera Assembler is a de novo Whole-Genome Shotgun (WGS) DNA sequence assembler.
GATK (Genome Analysis ToolKit) is a software package developed at the Broad Institute to analyze next-generation re-sequencing data.
HMMR is used for searching sequence databases for homologs of protein sequences, and for making protein sequence alignments. It implements methods using probabilistic models called profile Hidden Markov Models (HMMs).
BLAST is a bioinformatics algorithm for comparing primary biological sequence information, such as the amino-acid sequences of different proteins or the nucleotides of DNA sequences. A BLAST search enables a researcher to compare a query sequence with a library or database of sequences, and identify library sequences that resemble the query sequence above a certain threshold.
mpiBLAST is a freely available, open source, parallel implementation of NCBI BLAST.
NAMD is a parallel molecular dynamics code designed for high-performance simulation of large biomolecular systems
Picard comprises Java-based command-line utilities that manipulate SAM files, and a Java API (SAM-JDK) for creating new programs that read and write SAM files. Both SAM text and SAM binary (BAM) formats are supported.
SAMtools (Sequence Alignment/Map) tools format is a generic format for storing large nucleotide sequence alignments.
Velvet is a sequence assembler for very short reads.
VMD is a molecular visualization program for displaying, animating, and analyzing large biomolecular systems using 3-D graphics and built-in scripting.
General Science Applications & Libraries¶
CLHEP (Class Library for High Energy Physics) is a C++ library that provides utility classes for general numerical programming, vector arithmetic, geometry, pseudorandom number generation, and linear algebra, specifically targeted for high energy physics simulation and analysis software.
FDS (Fire Dynamics Simulator) is a Large-Eddy Simulation (LES) code for low-speed flows, with an emphasis on smoke and heat transport from fires.
SMV (SMokeView) is a visualization program used to display the output of FDS and CFAST simulations.
Geant4 is a toolkit for the simulation of the passage of particles through matter. Its areas of application include high energy, nuclear and accelerator physics, as well as studies in medical and space science.
GSL (GNU Scientific Library) is a numerical library for C and C++ programmers.
SciPy is a Python-based ecosystem of open source software for mathematics, science, and engineering.
NCAR Graphics is a Fortran and C based software package for scientific visualization.
netCDF & NCO (netCDF Operators) is a suite of programs designed to facilitate manipulation and analysis of self-describing data stored in the netCDF format.
OpenCV (Open source Computer Vision Library) is a library of programming functions mainly aimed at real-time computer vision.
PETSc (Portable, Extensible Toolkit for Scientific Computation) is a suite of data structures and routines developed by Argonne National Laboratory for the scalable (parallel) solution of scientific applications modeled by partial differential equations.
HDF5 (Hierarchical Data Format) is the name of a set of file formats (HDF/HDF4/HDF5) and libraries designed to store and organize large amounts of numerical data.
Trilinos is a collection of open source software libraries intended to be used as building blocks for the development of scientific applications.
NWChem is an open source high performance computational chemistry package.
GAMESS (General Atomic and Molecular Electronic Structure System) is a general ab initio quantum chemistry package.
PSI4 is an open source suite of ab initio quantum chemistry programs.
Mathematics and Statistics Applications & Libraries¶
R & Rmpi are a language and MPI library used for statistical computing and graphics.
MATLAB® is a high-level language and interactive environment for numerical computation, visualization, and programming.
NumPy is the fundamental package for scientific computing with Python.
ANN is an (Artificial Neural Network) software package for classification of remotely sensed data.
ATLAS (Automatically Tuned Linear Algebra Software) provides C and Fortran interfaces to an efficient BLAS implementation, as well as a few routines from LAPACK.
BLAS (Basic Linear Algebra Subroutine) is a de facto application programming interface standard for publishing libraries to perform basic linear algebra operations such as vector and matrix multiplication.
Boost is a set of C++ libraries that provide support for tasks and structures such as linear algebra, pseudorandom number generation, multithreading, image processing, regular expressions, and unit testing.
Intel® Math Kernel Libraries (MKL) include a wealth of routines to accelerate application performance and reduce development time.
FFTW (Fastest Fourier Transform in the West) is a software library for computing Discrete Fourier Transforms (DFTs).
ParMETIS is an MPI-based parallel library that implements a variety of algorithms for partitioning unstructured graphs, meshes, and for computing fill-reducing orderings of sparse matrices.
SPBLASTK is used for solving large sparse system of linear equations.
UDUNITS supports conversion of unit specifications between formatted and binary forms, arithmetic manipulation of units, and conversion of values between compatible scales of measurement.
Rendering, Remote Visualization and Image Manipulation¶
RealityServer is a software platform for the development and deployment of 3-D Web Services and 3-D applications.
JasPer is a project that creates a reference implementation of the codec specified in the JPEG-2000 Part-1 standard (i.e. ISO/IEC 15444-1). It consists of a C library and some sample applications useful for testing the codec.
ImageMagick is an open source software suite for displaying, converting, and editing raster image files. It can read and write over 200 image file formats.
Blender is a free and open source 3-D computer graphics software product used for creating animated films, visual effects, art, 3-D printed models, interactive 3-D applications and video games. Blender’s features include 3-D modeling, UV unwrapping, texturing, rigging and skinning, fluid and smoke simulation, particle simulation, soft body simulation, animating, match moving, camera tracking, rendering, video editing and compositing.
Compilers and Language Tools¶
Intel® C/C++ and Fortran compilers include optimization features and multithreading capabilities; highly optimized performance libraries; and error-checking, security, and profiling tools, allowing developers to create multithreaded applications and maximize application performance, security, and reliability.
Portland Group PGI® C/C++ and Fortran compilers incorporate global optimization, vectorization, software pipelining, and shared-memory parallelization capabilities.
CUDA® is a parallel computing platform and programming model developed by NVIDIA for general computing on Graphical Processing Units (GPUs). NVIDIA’s CUDA Compiler (NVCC) separates and sends host code (run on the CPU) to a C compiler, and GPU device code to the GPU which is further compiled by NVCC.
ANTLR (ANother Tool for Language Recognition) is a parser generator that uses
LL(*)parsing.FLTK (Fast, Light ToolKit) is a GUI library made to accommodate 3-D graphics programming.
GADL is a graphical programming project.
Gengetopt is a tool used to write command line option parsing code for C programs.
SWIG is a tool that easily allows a developer to wrap C/C++ functions for use with scripting languages.
Application Guides¶
WRF¶
Versions¶
There are various WRF versions available on POD. Use the module command to see the available versions. If you need any additional versions installed, please email support: pod@penguincomputing.com.
$ module avail wrf
----------------------------- /public/modulefiles ------------------------------
wrf/3.5/gcc.4.4.7 wrf/3.5/intel.12.1.0 wrf/3.5.1/intel.12.1.0
Example¶
A sample submission script is available at /public/examples/wrf-examples/wrf_example_1.sub. This is a self contained script that runs the default January 2000 case illustrated in the official WRF tutorial. To run the sample:
$ mkdir test_wrf
$ cp /public/examples/wrf-examples/wrf_example_1.sub test_wrf
$ cd test_wrf
$ qsub wrf_example_1.sub
To set up a working directory for running WPS + WRF + UPP from the public install, use the POD WRF utility script called build_wrf_workdir. This script takes a directory name as parameter and creates the appropriate folder structure for running WPS and WRF:
$ module load wrf/3.5.1/intel.12.1.0
$ mkdir test_wrf
$ build_wrf_workdir test_wrf
$ ls test_wrf
UPPV2.1 WPS WRFV3
Geogrid Data¶
Geogrid data for all available resolutions (30”, 2’, 5’ and 10’) are available in /public/apps/wrf/WPS_GEOG. The geog_data_path field of WRF’s namelist.wps has already been configured to use this geogrid data.
$ ls /public/apps/wrf/WPS_GEOG
albedo_ncep islope soiltype_bot_2m
greenfrac landuse_10m soiltype_bot_30s
hangl landuse_2m soiltype_bot_5m
hanis landuse_30s soiltype_top_10m
hasynw landuse_30s_with_lakes soiltype_top_2m
hasys landuse_5m soiltype_top_30s
hasysw maxsnowalb soiltype_top_5m
hasyw modis_landuse_20class_30s ssib_landuse_10m
hcnvx modis_landuse_21class_30s ssib_landuse_5m
hlennw orogwd_10m topo_10m
hlens orogwd_1deg topo_2m
hlensw orogwd_20m topo_30s
hlenw orogwd_2deg topo_5m
hslop orogwd_30m varsso
hstdv soiltemp_1deg
hzmax soiltype_bot_10m
MATLAB®¶
For MATLAB, users have the ability to submit batch jobs on POD. This means that their job is non-interactive and can be run completely at the command line. There is no access to the GUI. For instance, if a user had a MATLAB file named analyze.m on POD, they could run matlab -r analyze in their PBS script.
Example¶
Assuming the user has their MATLAB license information in $HOME/.matlab.lic and a MATLAB file named analyze.m, this is an example job submission script that can be submitted with the qsub command:
#PBS -S /bin/bash
#PBS -N MATLAB
#PBS -q H30
#PBS -l nodes=1:ppn=16
#PBS -j oe
module load matlab/R2013b
cd $PBS_O_WORKDIR
matlab -r analyze
exit $?
MATLAB® Distributed Computing Server (MDCS)¶
Users who have written parallel execution code using PCT (Parallel computing toolbox) can take advantage of MDCS on POD. They need to have a locally-installed MATLAB client, running on a Linux machine. They need to be licensed locally for MATLAB, PCT, and any additional toolboxes they want to use. With that local setup, the MATLAB GUI can be configured to submit PCT jobs to POD.
Please Note: These can only be batch jobs as interactive jobs are disabled because we do not allow TCP connections to be made between compute nodes and external IP addresses.
This documentation describes how to set up a MATLAB cluster configuration on your local client in order to run parallel jobs on POD using MDCS.
Licensing¶
In order to utilize MDCS on POD, the user must be licensed for MDCS as well. There are additional options here. We can access a local license (that you have provided to us and we host with Flex), or a remote license server (either publicly exposed, or via VPN). Email POD Support: pod@penguincomputing.com for assistance in configuring a VPN tunnel to a remote license server.
Additionally, the user may have access to MHLM (MathWorks Hosted License Manager). In this scenario, the user is paying for MDCS licenses in either a fixed amount (say, 32 workers for a year), or an on-demand amount (however many are checked out, up to some limit; for instance, 64 workers at a time). In the on-demand scenario, jobs are tracked to 1/10 minute (6 seconds), and all time is added up and billed at the end of the month.
When using MHLM, the client will be prompted for their MathWorks credentials when submitting the job. The use of MHLM is configured completely client-side and no work has to be done on POD to support it.
Initial Setup¶
For the initial setup file, please reference the documentation in a support ticket to pod@penguincomputing.com.
Once you have the MATLAB_POD.tar.gz file downloaded, extract its contents and enter the resultant directory:
$ tar -xzvf MATLAB_POD.tar.gz
$ cd MATLAB_POD
Before proceeding, have your MDCS license number handy if you will be using MathWorks Hosted License Management. You will also need to know the IP address of your POD login node and the username you use to login. Your IP(s) can be found on the main POD Portal page here.
Close down any MATLAB instances you have open, as the program only loads the Cluster Profile list when it starts up. Run the setup script, and follow its instructions.
$ ./setup_remote_cluster.sh
< M A T L A B (R) >
Copyright 1984-2013 The MathWorks, Inc.
R2013b (8.2.0.701) 64-bit (glnxa64)
August 13, 2013
To get started, type one of these: helpwin, helpdesk, or demo.
For product information, visit www.mathworks.com.
Depending on your MATLAB setup, either a dialog box will appear or you will be prompted the following questions:
Enter the address of your POD virtual machine: <Your POD Login Node IP>
Enter your POD user name: <Your POD Unix Username>
Are you using MathWorks Hosted License Management? [y/N] y
Enter your MHLM license number: <Your HMLM License Number>
The setup script will create a cluster profile named POD_remote_r2013b and will set it as your default cluster profile. In addition, the setup script will add the MATLAB_POD folder to your MATLAB search path.
Validation¶
As a quick check that the setup succeeded, you will need to validate it. Open up MATLAB, navigate to the Cluster Profile Manager (Parallel > Manage Cluster Profiles), select the POD_remote_r2013b profile, and click the Validate button. During this process, you will be asked to provide an identity file (your private SSH key used to log into POD) and whether the identity file is password protected. The answers will be saved and you will not need to enter them again.
The validation will run 5 tests. For security reasons, our firewall prevents starting hanging parallel pools on POD, so the final parpools test is expected to fail as POD does not support interactive parallel pools.
Customization¶
The setup script sets several default values that you may want to change. If you wish to modify the configuration, you may do so from the Cluster Profile Manager. Select the POD profile in the manager and click the Edit button to change any of the following parameters:
POD IP - If your POD IP changes, you may correct your configuration by modifying the second field in both the
independentSubmitFcnandcommunicatingSubmitFcntuples. You can always check your POD IP on the POD Portal page here.Worker Count - If you need to change the number of workers you use, you can do so by modifying the
NumWorkersentry. This value is set to 48 by the setup script.Remote Directory - If you need to use a different remote directory for job data storage, you may change it by editing the third field of both the
independentSubmitFcnandcommunicatingSubmitFcntuples. The default is your$HOME/MdcsDataLocation/POD/R2013b/remotedirectory on POD.Local Directory - If you need to change the local job data storage directory, you can do so by modifying the
JobStorageLocationentry.
Queue and Other Submission Options¶
The PBS job submission options can be controlled using the ClusterInfo class of functions provided by the MATLAB_POD environment.
Queue: The queue is initially set to
H30. If you wish to submit to a different queue, you may call theClusterInfo.setQueueName(<queue_name>)function. For instance:
>> ClusterInfo.setQueueName('M40')
PPN: The number of processors-per-node is initially set to 16, the appropriate value for the
H30queue. If you change the submission queue, remember to adjust the processors-per-node value using theClusterInfo.setProcsPerNode()function:
>> ClusterInfo.setQueueName('M40')
>> ClusterInfo.setProcsPerNode(12)
>> ClusterInfo.setQueueName('FREE')
>> ClusterInfo.setProcsPerNode(12)
Walltime: The wall clock time for the job can be set by the
ClusterInfo.SetWallTime('HH:MM:SS')function, for instance:
>> ClusterInfo.setWallTime('01:20:00')
Call the MATLAB function ClusterInfo.state() to display the current configuration:
>> ClusterInfo.state()
Arch :
ClusterHost : xxx.xxx.xxx.xxx
EmailAddress :
GpusPerNode :
MemUsage :
PrivateKeyFile : /home/user/.ssh/id_rsa
PrivateKeyFileHasPassPhrase : 1
ProcsPerNode : 16
ProjectName :
QueueName : H30
Reservation :
UseGpu : 0
UserDefinedOptions :
UserNameOnCluster : mypoduser
WallTime :
Please Note: Only some of the configuration options provided by the ClusterInfo class, for instance the GPU selection facilities, are not yet available. Thus activating these fields might result in jobs being rejected by the scheduler or jobs waiting forever in queue. At this stage, we recommend setting the QueueName, ProcsPerNode and WallTime properties.
LS-DYNA¶
Versions¶
There are multiple LS-DYNA versions installed on POD. Use the module command to see the available versions. If you need any additional versions installed, please email support: pod@penguincomputing.com.
$ module avail lsdyna
----------------------------- /public/modulefiles ------------------------------
lsdyna/10.0.0 lsdyna/9.0.1 lsdyna/dev_130556_s30
lsdyna/10.0.0_s30 lsdyna/9.1.0 lsdyna/dev_131124
lsdyna/10.1.0 lsdyna/9.2.0 lsdyna/dev_131137
lsdyna/10.1.0_s30 lsdyna/dev_115963 lsdyna/dev_131137_s30
lsdyna/11.0.0 lsdyna/dev_116406 lsdyna/dev_137250
lsdyna/11.0.0_s30 lsdyna/dev_116465 lsdyna/dev_138426
lsdyna/7.1.2 lsdyna/dev_120410
lsdyna/8.1.0 lsdyna/dev_130270
Example¶
More complex LS-DYNA examples with additional information about using LS-PREPOST can be found in /public/examples/lsdyna-examples. Below is a sample template for LS-DYNA:
#PBS -S /bin/bash
#PBS -q M40
#PBS -l nodes=4:ppn=12
#PBS -j oe
#PBS -N lsdyna
#PBS -r n
# LS-DYNA ARGUMENTS
LSDYNA_INPUT=neon.refined.rev01.k
# Work from the qsub folder
cd $PBS_O_WORKDIR
# Load LS-DYNA (Will load Intel and OpenMPI/Intel modules too)
module load lsdyna/7.0.0
# Execution
mpirun mpp971 I=$LSDYNA_INPUT
exit $?
LS-PREPOST Example¶
A sample template for LS-PREPOST:
#PBS -S /bin/bash
#PBS -q M40
#PBS -l nodes=1:ppn=12
#PBS -j oe
#PBS -N lsprepost
# Command file
CMD_FILE="neon_lspost.cfile"
# Work from the qsub folder
cd $PBS_O_WORKDIR
# Load lsprepost
module load lsprepost/4.1
# Execution (using Xvfb)
xvfb-run --server-args="-screen 0 1600x1200x24" lsprepost -nographics c="$CMD_FILE"
exit $?
LS-OPT¶
Versions¶
There are various LS-OPT versions available on POD. Use the module command to see the available versions. If you need any additional versions installed, please email support: pod@penguincomputing.com.
$ module avail lsopt
----------------------------- /public/modulefiles ------------------------------
lsopt/4.2 lsopt/5.0
Configuring¶
To configure LS-OPT to submit jobs to the PBS scheduler, one has to provide a custom script to submit the job, and tell LS-OPT that the jobs are executed by the scheduler. This is achieved by setting the appropriate keywords in the LS-OPT command file. For example:
...
solver dyna960 '1'
solver queue pbs
solver command "../../submit_smp_pbs"
solver concurrent jobs 0
...
In the above, the solver is LS-DYNA (solver dyna960 '1'), jobs are run by the PBS queue (solver queue pbs), and the solver command points to a script that submits the LS-DYNA job (sample submission scripts are shown in the Examples section). Note that the path to the submission script is specified relative to the working folder of the LS-DYNA job. This folder is created by LS-OPT at runtime, and the example assumes the submit_smp_pbs script is located in the same folder as the LS-OPT command file.
The last line (solver concurrent jobs 0) tells LS-OPT to submit all jobs in an iteration concurrently. This is the recommended setting when using a job scheduler.
Running¶
Once the LS-OPT command file and LS-DYNA submissions scripts are in place, LS-OPT can be run interactively on a login node. In this way the optimization calculation will be performed on the login node, while the LS-DYNA jobs will be executed on the cluster compute nodes.
To run LS-OPT on the login node type the following commands at the Linux prompt:
$ module load lsopt/4.2
$ lsopt <command-file>
Where <command-file> is the name of the LS-OPT command file
Examples¶
Sample files for the LS-OPT examples discussed below can be found in /public/examples/lsopt-examples. To run the sample calculation, copy the folder to your home storage then run LS-OPT interactively:
$ cp -ar /public/examples/lsopt-examples ~
$ cd ~/lsopt-examples
$ module load lsopt/4.2
$ lsopt run.opt
The run will submit a first batch of five LS-DYNA jobs for the first iteration, followed by a sixth job for the second iteration. The whole LS-OPT run will take about 5 minutes.
Using SMP LS-DYNA¶
Here is a template for using the SMP version of the LS-DYNA solver:
...
solver dyna960 '1'
solver queue pbs
solver command "../../submit_smp_pbs"
solver concurrent jobs 0
...
The submit_smp_pbs script is shown below:
#!/bin/bash
# Script to generate and submit an smp LS-DYNA job for an LS-OPT run
# Create a job name based on the working dir
jname=dynscr_$(pwd | sed -n 's/.*\/\(.*\)\/\(.*\)/\1\/\2/p')
cat > dynscr << EOF
#
# dynscr script
# =======================================================================
#PBS -S /bin/bash
#PBS -N ${jname}
#PBS -q H30
#PBS -l nodes=1:ppn=16
#
module load lsopt/4.2
module load lsdyna/7.1.1
export LSOPT_HOST=${LSOPT_HOST}
export LSOPT_PORT=${LSOPT_PORT}
cd \$PBS_O_WORKDIR
# The input file name is required for LS-OPT
wrapper smp971 ncpu=\$PBS_NP i=DynaOpt.inp
# =======================================================================
EOF
qsub dynscr
The above script submits the job to the H30 queue. To use the M40 queue two lines need to be changed:
#PBS -q M40
#PBS -l nodes=1:ppn=12
Using MPP LS-DYNA¶
For MPP LS-DYNA one needs a slightly different submission script:
...
solver dyna960 '1'
solver queue pbs
solver command "../../submit_mpp_pbs"
solver concurrent jobs 0
...
And the submit_mpp_pbs script is shown below:
#!/bin/bash
# Script to generate and submit an mpp LS-DYNA job for an LS-OPT run
# Create a job name based on the working dir
jname=dynscr_$(pwd | sed -n 's/.*\/\(.*\)\/\(.*\)/\1\/\2/p')
cat > dynscr << EOF
#
# dynscr script
# =======================================================================
#PBS -S /bin/bash
#PBS -N ${jname}
#PBS -q H30
#PBS -l nodes=1:ppn=16
#
module load lsopt/4.2
module load lsdyna/7.1.1
export LSOPT_HOST=${LSOPT_HOST}
export LSOPT_PORT=${LSOPT_PORT}
cd \$PBS_O_WORKDIR
# The input file name is required for LS-OPT
wrapper mpirun mpp971 i=DynaOpt.inp
# =======================================================================
EOF
qsub dynscr
The above script submits a 16 processors job to the H30 queue. One can modify the queue and node selection, for instance:
#PBS -q M40
#PBS -l nodes=4:ppn=12
StarCCM+¶
Versions¶
There are various StarCCM+ versions available on POD. Use the module command to see the available versions. If you need any additional versions installed, please email support: pod@penguincomputing.com.
$ module avail starccm
----------------------------- /public/modulefiles ------------------------------
starccm/7.02.011 starccm/8.02.008(default) starccm/8.04.010
starccm/7.04.011 starccm/8.02.011 starccm/8.06.005
starccm/7.06.012 starccm/8.04.007 starccm/8.06.007
Power on Demand License¶
To use your StarCCM+ Power on Demand license, choose the appropriate StarCCM+ version using modules and set $CMDLMD_LICENSE_FILE to use the CD-adapco license server. You will also need to define your project key as defined in your CD-adapco Power on Demand license configuration.
module load starccm/8.02.008
export CDLMD_LICENSE_FILE="1999@81.134.157.100"
export LM_PROJECT='enter_your_project_key_here'
Example¶
After configuring your license, leverage POD’s optimized OpenMPI with these commands:
module load openmpi/1.6.4/gcc.4.4.7
export OPENMPI_DIR=/public/apps/openmpi/1.6.4/gcc.4.4.7
You will then need to launch StarCCM+ with a number of flags to indicate the RSH binary to use, and the use of OpenMPI for MPI communication. Here is an example that can be found in /public/examples/starccm-examples:
#PBS -S /bin/bash
#PBS -N starccm
#PBS -l nodes=2:ppn=16
#PBS -j oe
#PBS -q H30
# environment for StarCCM+ using native OpenMPI
module load starccm/8.04.010
module load openmpi/1.6.4/gcc.4.4.7
export OPENMPI_DIR=/public/apps/openmpi/1.6.4/gcc.4.4.7
# set license environment for cd-adapco Power On Demand
export CDLMD_LICENSE_FILE='1999@flex.cd-adapco.com'
export LM_PROJECT='enter_your_project_key_here'
cd $PBS_O_WORKDIR
starccm+ -power -rsh /usr/bin/bprsh -batchsystem pbs -mpidriver openmpi -batch cavityQuad.java cavityQuad.sim
exit $?
ANSYS Fluent¶
Below is a template that can be used for ANSYS Fluent jobs on POD. Please contact support at pod@penguincomputing.com regarding license servers.
#PBS -S /bin/bash
#PBS -N fluent
#PBS -q M40
#PBS -l nodes=4:ppn=12
#PBS -l walltime=12:00:00
#PBS -j oe
# Load Fluent environment
module load fluent/14.5
export ANSYSLMD_LICENSE_FILE=<## YOUR LICENSE SERVER ##>
export ANSYSLI_SERVERS=<## YOUR LICENSE SERVER ##>
cd ${PBS_O_WORKDIR}
nprocs=$(wc -l $PBS_NODEFILE | awk '{print $1}')
fluent 3ddp -g -t${nprocs} -cnf=${PBS_NODEFILE} -i <## INPUT FILE ##>
exit $?
ANSYS Electronics Desktop¶
How to configure ANSYS Electronics Desktop to submit to POD Queues
Open ANSYS Electronics Desktop.
Click Tools -> Job Management -> Select Scheduler.
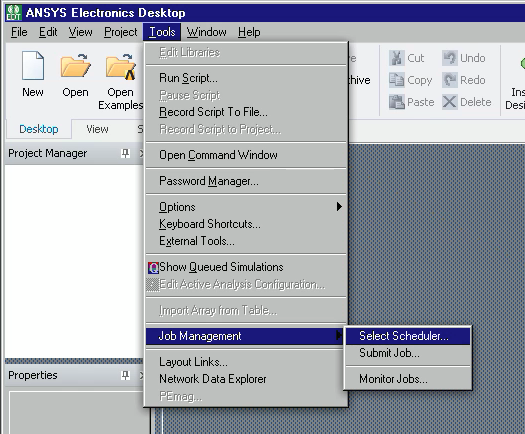
Choose “pbs” from “Choose scheduler”, then click Refresh button, then OK.
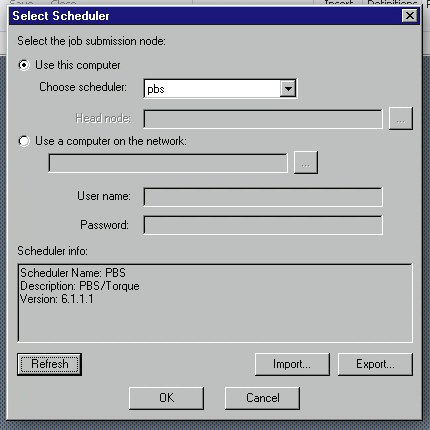
How to submit a job from ANSYS Electronics Desktop to POD Queue
Click Tools -> Job Management -> Submit Job.
Browse to project via “Project Path”.
Under “Compute Resources” tab, click the “…” button next to “Resource selection parameters”.
Check “Queue” and choose desired Queue under the Value column, then click OK button.
Uncheck “Use automatic settings”.
Specify “Total number of task” (number of nodes) and “Cores per distributed task” (cores per node).
The job name can be set under the Scheduler Options tab.
Click Submit Job button.
R & Rmpi¶
Various versions of R are available on POD using the module command. If you need any additional versions installed, please email pod@penguincomputing.com.
$ module avail R
----------------------------- /public/modulefiles ------------------------------
R/3.0.0/gcc.4.4.7(default) Rmpi/0.6-3/R.3.0.0
R/3.0.2/gcc.4.4.7
Rmpi¶
Rmpi is available by loading the corresponding environment module. The module load command will also load any required dependencies modules automatically:
$ module load Rmpi/0.6-3/R.3.0.0
$ module list
Currently Loaded Modulefiles:
1) gcc/4.4.7 3) openmpi/1.6.4/gcc.4.4.7
2) R/3.0.0/gcc.4.4.7 4) Rmpi/0.6-3/R.3.0.0
Running Rmpi Programs¶
Rmpi programs are typically executed by starting a master task which spawns the MPI slave processes. The master task has to be started using the mpirun command in order to initialize the MPI environment, however only 1 MPI process need to be started at this time, since the slave processes are spawned internally by the master task. Thus the typical command used to start an Rmpi program would be the following:
$ mpirun -np 1 R --slave CMD BATCH <my-Rmpi-program.R>
Spawning MPI Processes¶
The Rmpi function mpi.spawn.Rslaves() is used to spawn MPI tasks. This function takes an optional argument specifying how many slaves have to be spawned. The following snippet of code can be used to start as many task as the MPI environment allows. This is achieved by polling the environment variable $OMPI_UNIVERSE_SIZE and subtracting 1 to account for the master task:
# Spawn as many slaves as possible
NS <- type.convert(Sys.getenv("OMPI_UNIVERSE_SIZE")) - 1
mpi.spawn.Rslaves(nslaves=NS)
Rmpi Examples¶
Rmpi examples and submission scripts are available at /public/Rmpi-examples. Here is an example:
#PBS -S /bin/bash
#PBS -q M40
#PBS -N Rmpi
#PBS -j oe
#PBS -l nodes=4:ppn=12
module load Rmpi
cd $PBS_O_WORKDIR
echo "Job ID: $PBS_JOBID"
echo "Queue: $PBS_QUEUE"
echo "Cores: $PBS_NP"
echo "Nodes: $(cat $PBS_NODEFILE | sort -u | tr '\n' ' ')"
echo "mpirun: $(which mpirun)"
echo "R: $(which R)"
mpirun -np 1 R --slave CMD BATCH example.R
exit $?
# vim: syntax=sh
Where, example.R reads:
# Load the R MPI package if it is not already loaded.
if (!is.loaded("mpi_initialize")) {
library("Rmpi")
}
# Spawn as many slaves as possible
NS <- type.convert(Sys.getenv("OMPI_UNIVERSE_SIZE")) - 1
mpi.spawn.Rslaves(nslaves=NS)
# In case R exits unexpectedly, have it automatically clean up
# resources taken up by Rmpi (slaves, memory, etc...)
.Last <- function(){
if (is.loaded("mpi_initialize")){
if (mpi.comm.size(1) > 0){
print("Please use mpi.close.Rslaves() to close slaves.")
mpi.close.Rslaves()
}
print("Please use mpi.quit() to quit R")
.Call("mpi_finalize")
}
}
# Tell all slaves to return a message identifying themselves
mpi.remote.exec(paste("I am",mpi.comm.rank(),"of",mpi.comm.size()))
# Tell all slaves to close down, and exit the program
mpi.close.Rslaves()
mpi.quit()
Singularity¶
POD supports Singularity, a powerful Linux container platform designed for High Performance Computing. Singularity enables users to have full control of their environment, enabling a non-privileged user to swap out the operating system on the host for one they control. As an example, Singularity can provide a user with the ability to create an Ubuntu image of their application, and run the containerized application on POD in its native Ubuntu environment. For more information:
Singularity Home: https://sylabs.io/docs/
Google Groups: https://groups.google.com/a/lbl.gov/forum/#!forum/singularity
Creating Singularity Containers¶
To use Singularity on POD one needs to either create a Singularity container or use one provided by someone else. To build a container, one needs root access on the build system so one cannot build one on POD. Some options for a build environment are:
If you have root access to a Linux system you can install Singularity and build containers there.
If you don’t have a Linux system you could easily install one in a virtual machine using software like VirtualBox, VMware, or Vagrant.
You can create containers and download them from here or Vagrant Singularity Boxes for pre-built Singularity environments (if using Vagrant as your virutal machine).
Once you have built and tested your container locally, just use SCP or Rsync to upload it to your POD storage.
$ scp -C my_container.img [login node address]:~/
or
$ rsync -z my_container.img [login node address]:~/
Using Docker Images in Singularity¶
Docker images can be used to create Singularity or can even be run directly. More information can be found at Singularity’s home page, but we’ve used two different methods to incorporate Docker into Singularity on POD.
Method 1 - Importing a Docker image into a Singularity container
Singularity communicates with Docker Hub using the Docker Remote API. One can create a Singularity image and import from a docker ::
Please Note: That this is a one way operation. Changes made to the image will not be propagated to the Docker image.
remote$ sudo singularity build tensorflow.img docker://tensorflow/tensorflow:latest
Method 2 - Using a Singularity Recipe file to build from Docker Hub
In the recipe file, specify docker as the Bootstrap source and specify the image in Docker syntax
Bootstrap: docker
From: tensorflow/tensorflow:latest
Singularity Recipes created by Penguin for POD¶
While running single node containers on POD is very easy, creating a container for MPI is a bit more complex. Penguin has created several skeleton recipe files as examples of creating images suitable for running on POD. You can find them in the PenguinComputing/POD public git repository.
There you will find example recipe files for CentOS and Ubuntu with OpenMPI using various network transports supported on POD and a script to make creating and bootstrapping containers easier. Check back for additional spec files and for special requests please reach out to POD support team: pod@penguincomputing.com.
Here is an example:
# copy pod repo to your local machine
remote$ git clone https://github.com/PenguinComputing/pod.git
remote$ cd pod/singularity
# if you don’t have git installed on your local machine you can also
# browse to https://github.com/PenguinComputing/pod.git and download a spec file
# build the pod centos7 recipe (.def) into a sandbox folder
remote$ sudo singularity build --sandbox ./pod-ompi2-centos7 ./pod-ompi2-centos7.def
# convert the sandbox into a read-only, squashfs image
remote$ sudo singularity build ./pod-ompi2-centos7.img ./pod-ompi2-centos7
# copy the final image to pod; compressed transfers are recommended (rsync -z)
remote$ rsync -azvH ./pod-centos7-ompi2.img [pod username]@[login node address]:~/
# login into pod, and submit a job in the B30 queue; this example uses interactive qsub
remote$ ssh [pod username]@[login node address]
pod$ qsub -I -q B30 -l nodes=2:ppn=28,walltime=00:15:00
# in your job environment, load singularity and ompi 2.0.1
n550$ module load singularity/2.4
n550$ module load openmpi/2.0.1/gcc.4.8.5
# run the mpi_ring application inside the container using pod's mpirun
n550$ mpirun singularity exec pod-ompi2-centos7.img /usr/bin/mpi_ring
Tips for running MPI applications with Singularity on POD¶
Always compile MPI applications inside a container image with the same MPI implementation and version you plan to use on POD. Refer to the Singularity documentation for currently supported MPI implementations.
The POD B30 and S30 queues are enabled by Intel’s Omni-Path fabric, which works optimally with the PSM2 transport for MPI communication. Not all Operating Systems support PSM2. CentOS 7 and Ubuntu’s Beta Zesy release (17.04) can be built with PSM2 support. Ubuntu 16 does not support PSM2 and will require the use of IPoIB for MPI communication.
ClamAV® Antivirus¶
POD provides access to the ClamAV open source antivirus engine for detecting trojans, viruses, malware and other malicious threats. Users can scan files, directories and even whole directory trees on demand. More information can be found on the ClamAV website.
POD administrators keep the virus database up to date so all users have to do is scan any files or directories that they wish.
ClamAV may flag harmless files as infected. If you find any infected files, please notify POD support: pod@penguincomputing.com. We can check to see if you indeed have an infected file or if ClamAV has generated a false positive.
Scanning¶
Here is an example of scanning the current working directory. This example uses clamscan -i to only print the names of infected files rather than all files it scans.
$ module load clamav
$ clamscan -i
----------- SCAN SUMMARY -----------
Known viruses: 6298901
Engine version: 0.99.2
Scanned directories: 1
Scanned files: 39
Infected files: 0
Data scanned: 1.27 MB
Data read: 1.00 MB (ratio 1.27:1)
Time: 13.835 sec (0 m 13 s)
To scan a single file, just tell clamscan which file to scan:
$ module load clamav
$ clamscan Makefile
Makefile: OK
----------- SCAN SUMMARY -----------
Known viruses: 6298901
Engine version: 0.99.2
Scanned directories: 0
Scanned files: 1
Infected files: 0
Data scanned: 0.03 MB
Data read: 0.02 MB (ratio 2.00:1)
Time: 13.147 sec (0 m 13 s)
To scan all files and subdirectories in your home directory
$ module load clamav
$ clamscan -r $HOME
To lean more about clamscan, please visit the ClamAV website or read the manual page on POD.
$ man clamscan